OKA LABORATORY
2026
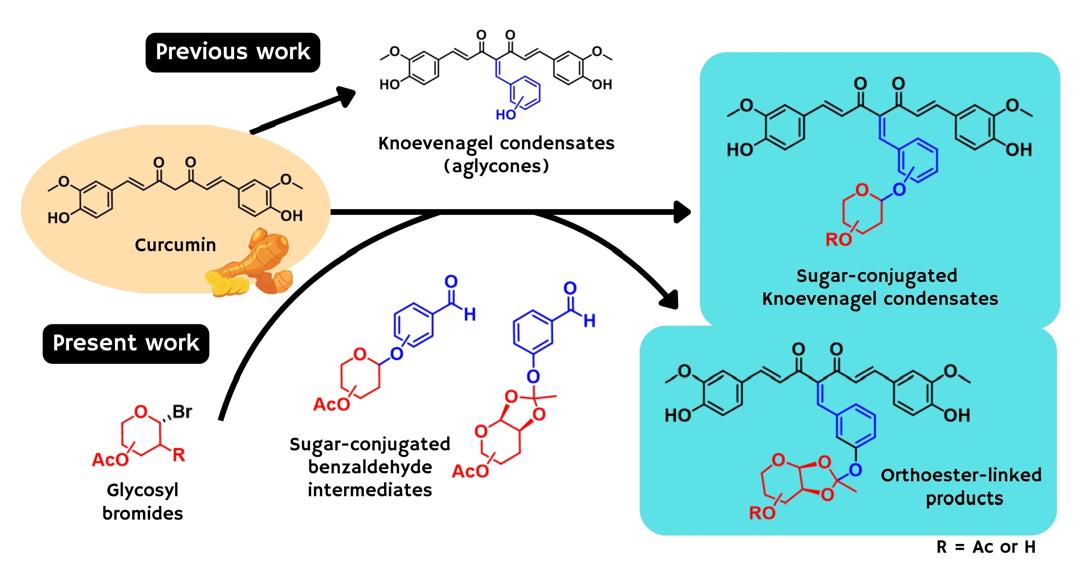
60. Novel sugar-conjugated Knoevenagel condensate curcumin derivatives as promising bioactive hybrids with antimalarial potential. Jamil, S. N. H.; Oka, N.; Ali, A. H.; Ng, Y. H.; Marzuki, N. F. N.; Feroz, S. R.; Lam, S. D.; Mahmud, F.; Zakaria, Y.; Latip, J. RSC Adv. 2026, 16, 24689-24703.
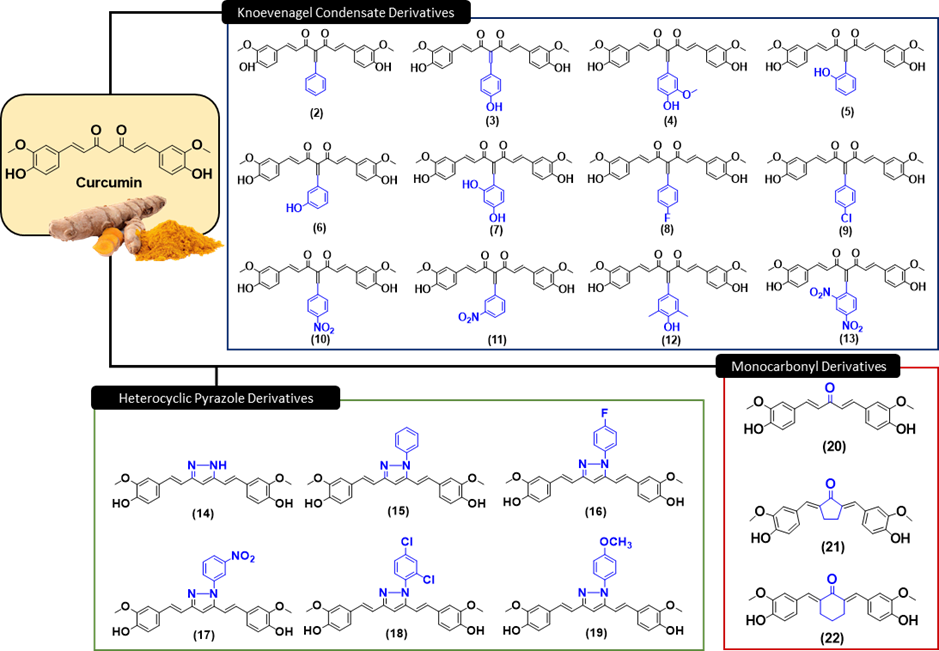
59. Antimalarial potential of curcumin derivatives evaluated through experimental and computational approaches. Jamil, S. N. H.; Bakar, K. A.; Ali, A. H.; Marzuki, N. F. N.; Shahid, F.; Mahmud, F.; Feroz, S. R.; Lam, S. D.; Oka, N.; Zakaria, Y.; Muhajir, M. I.; Maharani, R.; Supratman, U.; Latip, J. Sci. Rep. 2026, 16, 2512.
2025
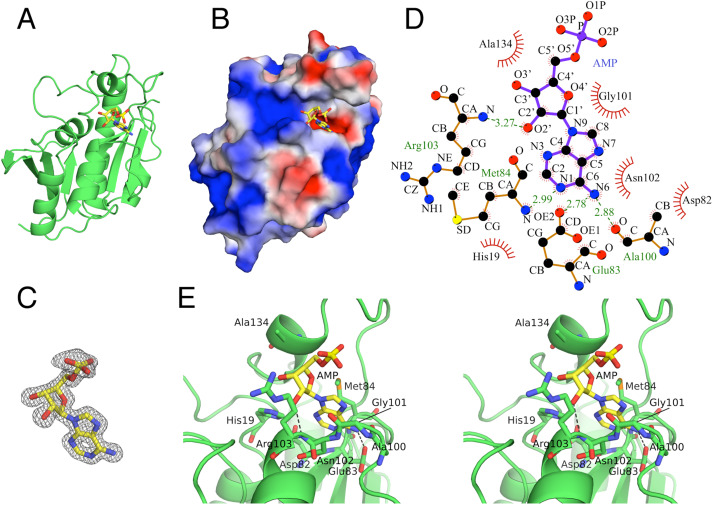
55. Binding mode between peptidyl-tRNA hydrolase and the peptidyl-A76 moiety of the substrate. Uehara, Y.; Matsumoto, A.; Nakazawa, T.; Fukuta, A.; Ando, K.; Uchiumi, T.; Oka, N.; Ito, K. J. Biol. Chem. 2025, 301, 108385.

54. The Protective Effects of Carvacrol Against Doxorubicin-Induced Cardiotoxicity In Vitro and In Vivo. Retnosari, R.; Ghani, M. A. A.; Alkharji, M. M.; Nawi, W. N. I. S. W.; Rushdan, A. S. A.; Mahadi, M. K.; Ugusman, A.; Oka, N.; Zainalabidin, S.; Latip, J. Cardiovasc. Toxicol. 2025, 25, 167-181.
2024
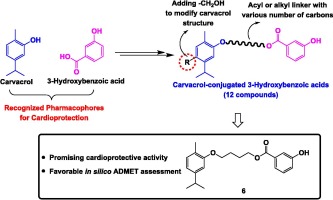
53. Carvacrol-conjugated 3-hydroxybenzoic acids: design, synthesis, cardioprotective potential against doxorubicin-induced cardiotoxicity, and ADMET study. Retnosari, R.; Oh-hashi, K.; Ugusman, A.; Zainalabidin, S.; Latip, J.; Oka, N. Bioorg. Med. Chem. Lett. 2024, 113, 129973.
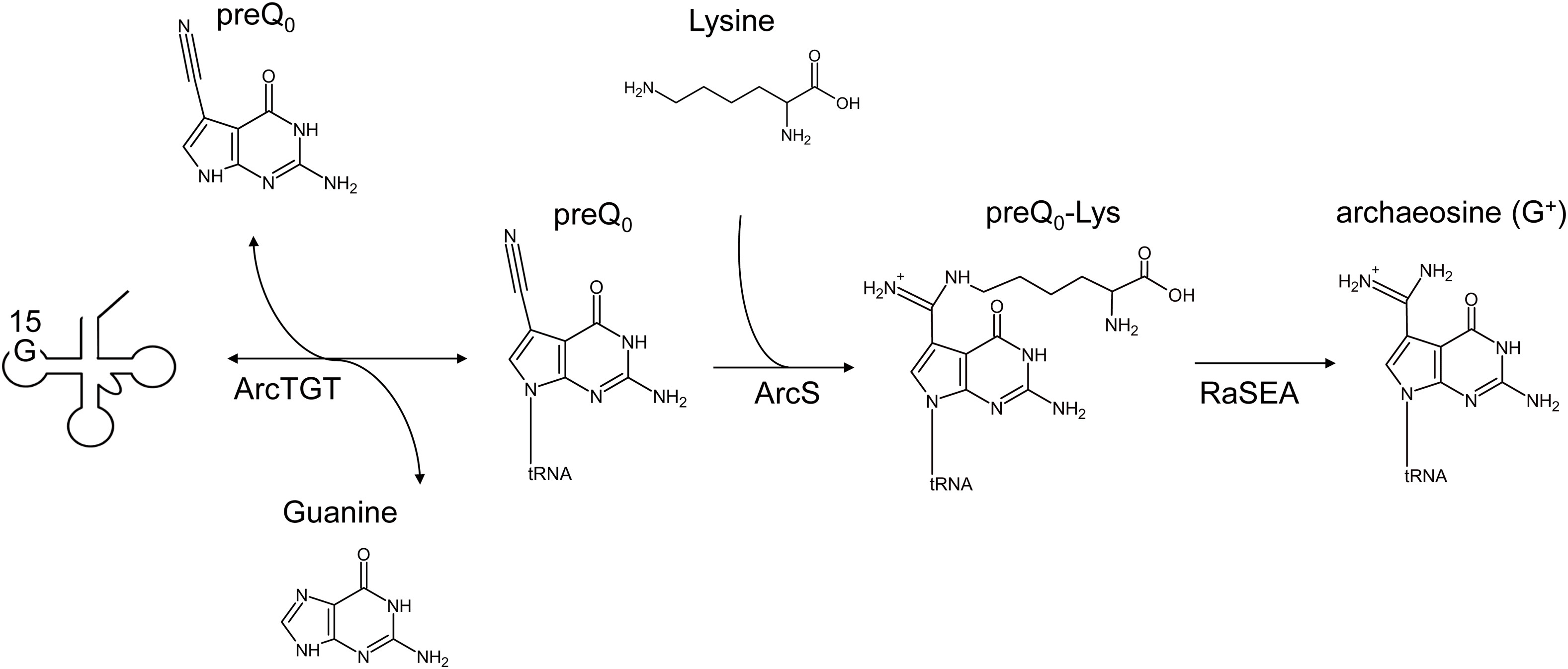
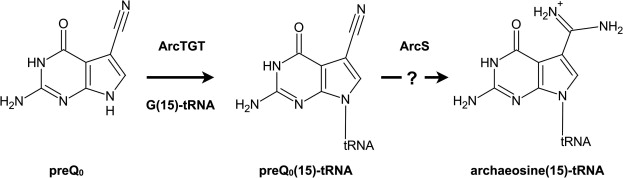
52. ArcS from Thermococcus kodakarensis transfers L-lysine to preQ0 nucleoside derivatives as minimum substrate RNAs. Fujita, S.; Sugio, Y.; Kawamura, T.; Yamagami, R.; Oka, N.; Hirata, A.; Yokogawa, T.; Hori, H. J. Biol. Chem. 2024, 300, 107505.
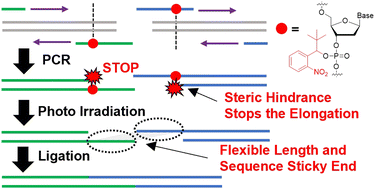
51. Development of PCR primer enabling the design of flexible sticky ends for efficient concatenation of long DNA fragments. Nomura, K.; Onda, K.; Murase, H.; Hashiya, F.; Ono, Y.; Terai, G.; Oka, N.; Asai, K.; Suzuki, D.; Takahashi, N.; Hiraoka, H.; Inagaki, M.; Kimura, Y.; Shimizu, Y.; Abe N.; Abe, H. RSC Chem. Biol. 2024, 5, 360-371.
2023
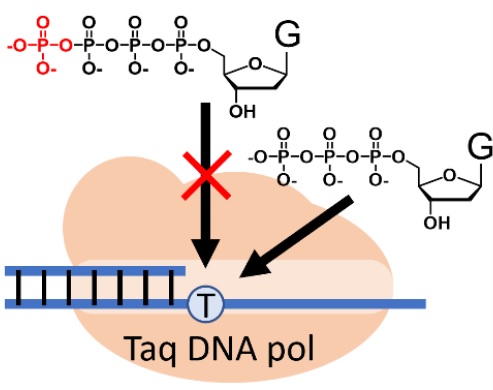
50. The effect of γ phosphate modified deoxynucleotide substrates on PCR activity and fidelity. Hashiya, F.; Murase, H.; Chandela, A.; Hiraoka, H.; Inagaki, M.; Nakashima, Y.; Abe, N.; Nakamura, M.; Terai, G.; Kimura, Y.; Ando, K.; Oka, N.; Asai, K.; Abe, H. ChemBioChem 2023, 24, e202200572.
2021
~2020
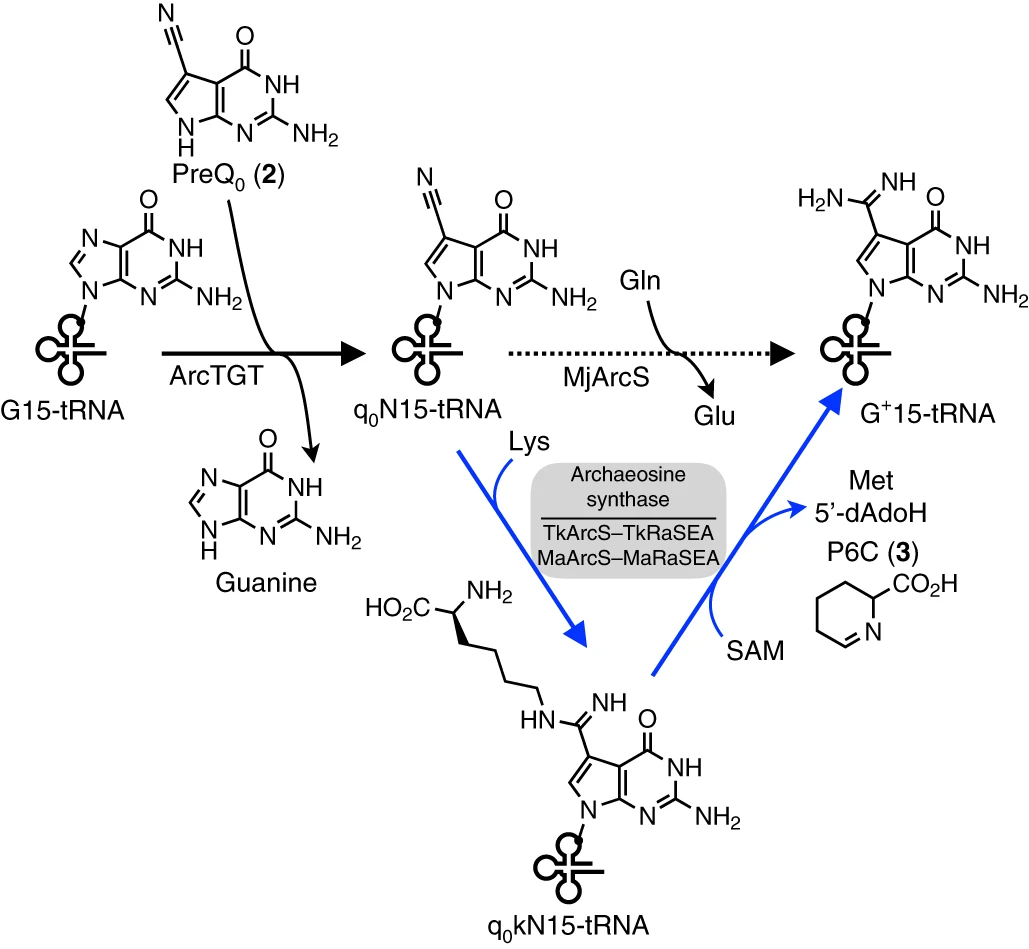
44. Identification of a radical SAM enzyme involved in the synthesis of archaeosine. Yokogawa, T.; Nomura, Y.; Yasuda, A.; Ogino, H.; Hiura, K.; Nakada, S.; Oka, N.; Ando, K.; Kawamura, T.; Hirata, A.; Hori, H.; Ohno, S. Nat. Chem. Biol. 2019, 15, 1148-1155.
32. Purification and comparison of native and recombinant tRNA-guanine transglycosylases from Methanosarcina acetivorans. Nomura, Y.; Onda, Y.; Ohno, S.; Taniguchi, H.; Ando, K.; Oka, N.; Nishikawa, K.; Yokogawa, T. Protein Expr. Purif. 2013, 88, 13-19.
↑